This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
Protein Interaction Networks
|
What are protein interaction networks?
Protein interaction networks depict the connections between many proteins in a specific organism. In some cases, like in the image to the right, there are a lot of interactions that are shown in a web-pattern. In most cases, the interactions include control, suppression, and activation, among other things. Although it can be hard to discern images like this, looking at specific proteins can narrow the search and lead to a much more manageable web. |
Image 1. Protein interaction network example. Image credit
|
Databases like STRING, IntAct, and BioGrid show known and predicted protein interactions from scientific research. Each database connects the proteins through string-type models and shows their individual function in cells. Using this information, it can be hypothesized how a protein works in the cell. The function of the proteins that interact with the protein of interest give a more in-depth look into processes controlled by the gene of interest.
Below are the images generated by the databases. After looking at the complex string maps, the proteins were separated based on function and location. This gives researchers, like myself, more insight into how the protein interactions lead to phenotypes in the body.
Below are the images generated by the databases. After looking at the complex string maps, the proteins were separated based on function and location. This gives researchers, like myself, more insight into how the protein interactions lead to phenotypes in the body.
Discussion:
In the case of the MAPT gene (the Tau protein), MAPT is connected to proteins associated with mitosis, meiosis, the cell cycle, cell differentiation, and apoptosis (controlled cell death). These connections make sense as MAPT is also known to control the cell cycle and lead to cell differentiation in the brain.
In the case of the MAPT gene (the Tau protein), MAPT is connected to proteins associated with mitosis, meiosis, the cell cycle, cell differentiation, and apoptosis (controlled cell death). These connections make sense as MAPT is also known to control the cell cycle and lead to cell differentiation in the brain.
Proteomics
What are proteomics?
Proteomics studies proteins, their modification, function, and interactions within biology. Several databases were used to analyze the protein and answer several important questions used in research. Using this information can give insight into how the protein works and how mutations might affect the protein and its function. The databases used include Expasy, NetPhos, and Locate.
Proteomics studies proteins, their modification, function, and interactions within biology. Several databases were used to analyze the protein and answer several important questions used in research. Using this information can give insight into how the protein works and how mutations might affect the protein and its function. The databases used include Expasy, NetPhos, and Locate.
What questions can be answered?
1. What enzymes cut the protein?
Many enzymes cleave the protein at specified locations. Some of the more notable enzymes include Pepsin, Proteinase K, and Trypsin. Each of these enzymes cut the Tau protein multiple times throughout the amino acid sequence, which can lead to altered protein expression and function.
Many enzymes cleave the protein at specified locations. Some of the more notable enzymes include Pepsin, Proteinase K, and Trypsin. Each of these enzymes cut the Tau protein multiple times throughout the amino acid sequence, which can lead to altered protein expression and function.
2. What is the isoelectric point of the protein?
The isoelectric point of the protein is the pH at which the protein has no electrical charge. For the Tau protein, it is a pH of 6.61.
The isoelectric point of the protein is the pH at which the protein has no electrical charge. For the Tau protein, it is a pH of 6.61.
3. What is the molecular weight of the protein?
The Tau protein has a molecular weight of 80,940.87 grams per mole.
The Tau protein has a molecular weight of 80,940.87 grams per mole.
4. Where does the protein localize?
According to the Locate database, the Tau protein localizes in the nucleus of the cell.
According to the Locate database, the Tau protein localizes in the nucleus of the cell.
5. What post-translational modifications are made to the protein?
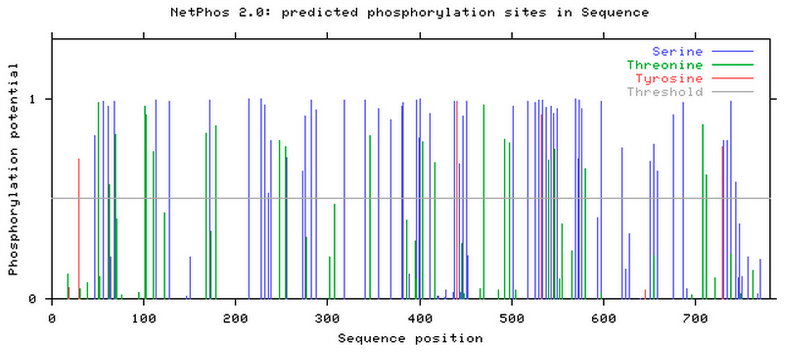
The Tau protein has multiple phosphorylation sites, including 57 serine sites, 21 threonine sites, and 4 tyrosine sites. These are considered post-modification sites because the added phosphorylase happens after transcription.
The Tau protein has multiple phosphorylation sites, including 57 serine sites, 21 threonine sites, and 4 tyrosine sites. These are considered post-modification sites because the added phosphorylase happens after transcription.
The phosphorylation sites for the Tau protein, according to NetPhos.
6. How well conserved are these modifications across species?
Four of the phosphorylation sites are in the same region as the previously studied domains in the Tau protein. These sites are well conserved across species such as primates, rodents, cats, and dogs.
Four of the phosphorylation sites are in the same region as the previously studied domains in the Tau protein. These sites are well conserved across species such as primates, rodents, cats, and dogs.
Discussion
Using the Proteomics databases, the information gleaned can further research. In some cases, phosphorylation sites occur at the same areas as protein domains. For the Tau protein, this is the case. Therefore, further research needs to be done to determine if phosphorylation is having an influence on Tau protein levels and, to a further extent, FTD. Analysis on induced or suppressed phosphorylation could lead to more information or even a treatment.
Using the Proteomics databases, the information gleaned can further research. In some cases, phosphorylation sites occur at the same areas as protein domains. For the Tau protein, this is the case. Therefore, further research needs to be done to determine if phosphorylation is having an influence on Tau protein levels and, to a further extent, FTD. Analysis on induced or suppressed phosphorylation could lead to more information or even a treatment.
Site built by Kassandra Ford
Genetics 564, Spring 2015
University of Wisconsin-Madison
Site last updated: 5/13/2015
Genetics 564, Spring 2015
University of Wisconsin-Madison
Site last updated: 5/13/2015