This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
Protein Domains and Motifs
|
Protein domains are regions of the gene, in this case MAPT, that are exceptionally conserved across species. Although a function may not be known, scientists use domains to predict areas that are imperative to gene function.
When comparing domains, it is standard to use multiple software programs to compare amino acid sequences that encode for proteins. In our research, there were several programs used for different purposes. PFAM, SMART, InterPro, and PROSITE were used to predict the domains of this Tau protein. The results are listed on the right. All four software programs predicted this four-block domain shape seen in all organisms. GO and UniProt programs were used to predict the function of these domains (see Gene Ontology section). Programs such as PFAM and SMART are able to depict projections for what the protein sequence looks like, in terms of exons, introns, and important domains. PFAM also has the capabilities to generate circular trees with all species known to have the protein sequence. Images from these two program are shown below under the associated program name. DNA Motifs are short, repeated sequences of DNA that are assumed to have important function. Similar to protein motifs, further research is needed to determine if these sequences are truly critical, but if the sequences are conserved across many species, it is likely that there is a use for them. |
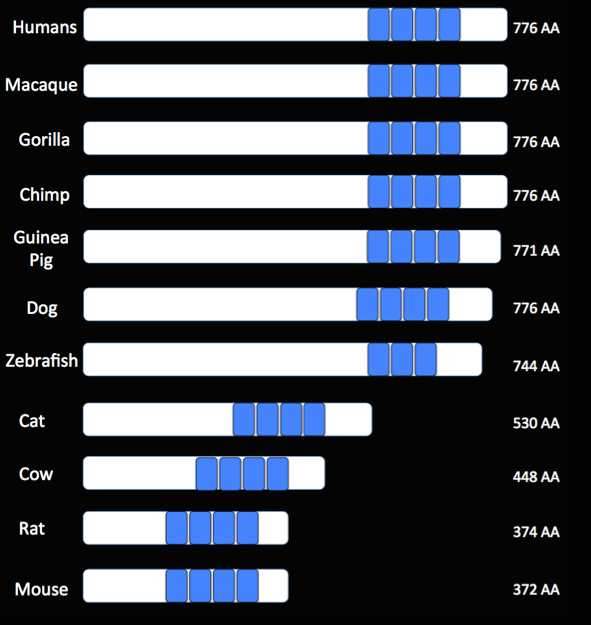
Figure 1. An image generated to visualize the relative position and conservation of the protein domains across multiple organisms. Nearly all species contained the four domains, with one exception. The relative size of the Tau protein sequences can also be compared across species.
|
What domains were found?
Four domains were found in the MAPT gene, all four being tubulin-binding domains. The four domains are all directly next to each other, with introns separating the domains prior to mRNA processing. Because the domains are directly related to the exon splicing process, correct splicing is hypothesized to be imperative to MAPT function.
Four domains were found in the MAPT gene, all four being tubulin-binding domains. The four domains are all directly next to each other, with introns separating the domains prior to mRNA processing. Because the domains are directly related to the exon splicing process, correct splicing is hypothesized to be imperative to MAPT function.
SMART Images
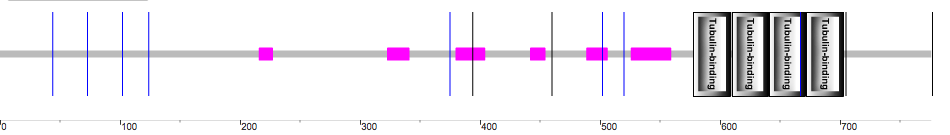
Figure 2. The SMART depiction of the MAPT gene and the four tubulin-binding domains.
Where do the mutations generally occur?
As stated previously, many mutations have been found in the MAPT gene. The most known mutations are in exon 10, with quite a few additional mutations known in intron 10 and exon 12. Those mutations are shown in the figure below. When the location of the protein domains is considered, it would be hypothesized that mutations in areas like exon 10 and intron 10 (closest to the domains) would have the greatest effect on protein function. Altering the gene domains (the most important parts of the gene) could have serious implications for protein function and expression.
As stated previously, many mutations have been found in the MAPT gene. The most known mutations are in exon 10, with quite a few additional mutations known in intron 10 and exon 12. Those mutations are shown in the figure below. When the location of the protein domains is considered, it would be hypothesized that mutations in areas like exon 10 and intron 10 (closest to the domains) would have the greatest effect on protein function. Altering the gene domains (the most important parts of the gene) could have serious implications for protein function and expression.
Figure 3. The positions of exons and introns in MAPT with the respective number of mutations found in each location. Above image credit
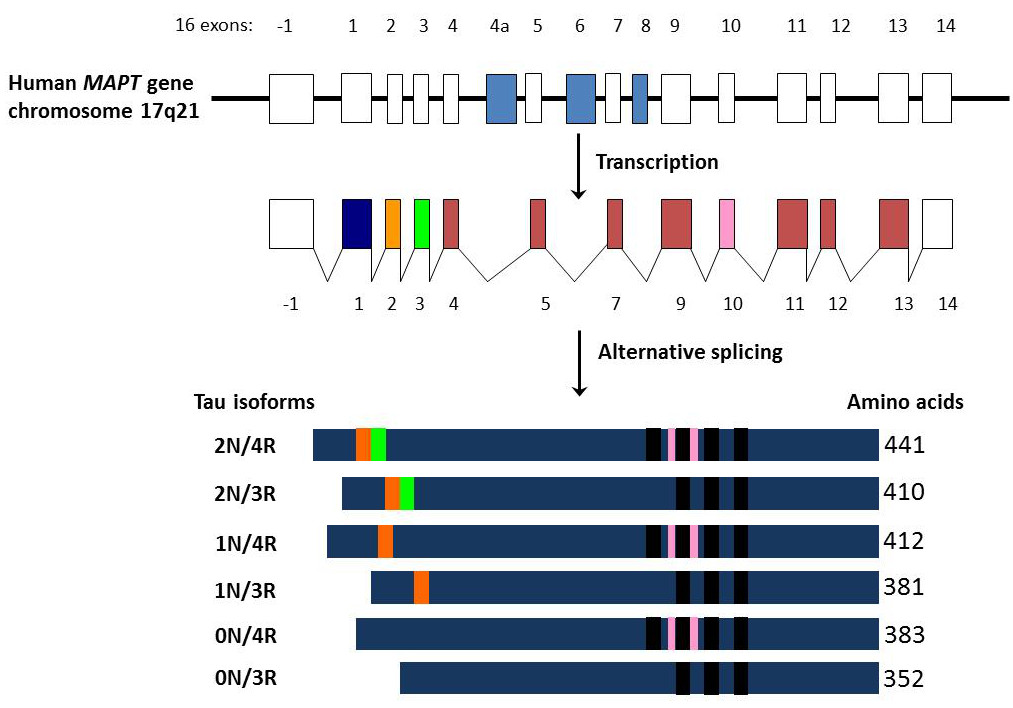
Figure 4. The relative exons and introns of MAPT, along with the transcription and alternative splicing processes. The mutations in exon 10 directly surround one of the protein domains seen above. Below image credit
Figure 4. The relative exons and introns of MAPT, along with the transcription and alternative splicing processes. The mutations in exon 10 directly surround one of the protein domains seen above. Below image credit
When the location of the domains is compared to the location of the mutations, it can be hypothesized that there may be a connection between the location of the mutations and altered protein levels seen in the FTD phenotype. This could be a molecular reason behind FTD and its phenotype. Further research about the protein domains is proposed in the Specific Aims portion of the website.
PFAM Images
Where do the proteins localize?
Although Tau proteins are located throughout the body and are important for cell differentiation and proliferation, the focus of this study is on the Tau proteins located within the nervous system and the brain. The proteins are important in neuron function and play pivotal roles in neural functioning.
Although Tau proteins are located throughout the body and are important for cell differentiation and proliferation, the focus of this study is on the Tau proteins located within the nervous system and the brain. The proteins are important in neuron function and play pivotal roles in neural functioning.
Discussion
After finding the domains of the MAPT gene and determining the purpose of those domains, it can be hypothesized that mRNA processing is important for obtaining correct MAPT expression and protein formation. Mutations in exon and intron 10 could have profound implications on the correct processing of the domains. These locations are also known to have the most mutations. Further research should be done to determine the molecular effects of mutations in these two locations.
After finding the domains of the MAPT gene and determining the purpose of those domains, it can be hypothesized that mRNA processing is important for obtaining correct MAPT expression and protein formation. Mutations in exon and intron 10 could have profound implications on the correct processing of the domains. These locations are also known to have the most mutations. Further research should be done to determine the molecular effects of mutations in these two locations.
Chemical Genetics
|
What is Chemical Genetics?
Chemical genetics is the use of biotechnology to alter protein function. The process involves creating different "small molecules" from a basic structure. A screening process is then used to determine if any of these molecules could be important to a particular protein's function. This chemical screening involves a microarray. Either the small molecule is placed on the microarray surface and the proteins are flushed down the surface, or vis versa. The goal is to have the small molecule bind to the protein. During this process a fluorescent light will be shown if the protein and the small molecule have an exact match [1]. Once a match is determined, the small molecule is isolated and tested further. There are several ways to organize the small molecules: diversity oriented and target-oriented libraries, based on the library's purpose [1]. Diversity oriented libraries accumulate a large variety of small molecules, based on a few similar structures. Target oriented libraries are based off one structure and have a few very small changes. |
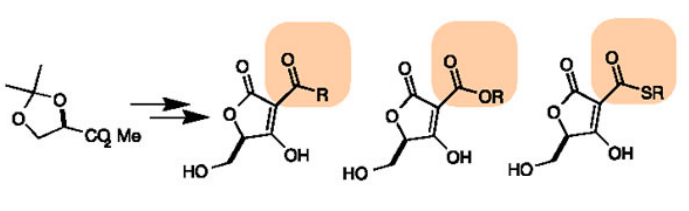
Figure 5. An example of a diversity-oriented library Image credit
Figure 6. An example of a target-oriented library. Image credit
|
Chemical genetics in FTD
|
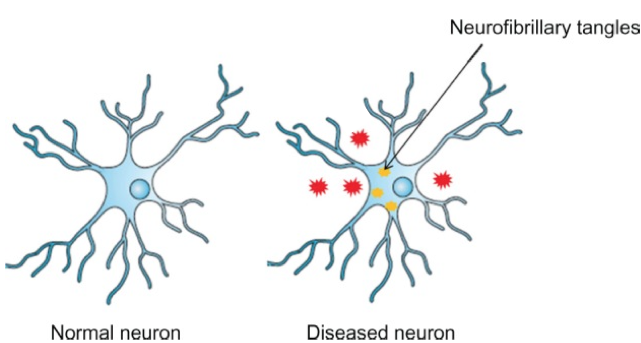
Chemical Genetics databases like PubChem and ChemBank exist to aid researchers in finding information about particular proteins, phenotypes, or genes. After a search for FTD and the gene/protein associated with the disease, no results were found. Small molecules that are associated with Alzheimer's were the only ones present. When diseases like Alzheimer's and FTD are present, tau proteins are unable to bind to microtubules and instead aggregate. These aggregations, called neurofibrillary tangles, can be recognized and treated with various drugs. Below are a list of drugs that can bind to and alter the function of the neurofibrillary tangles.
|
Figure 7. A normal neuron cell compared to one that contains neurofibrillary tangles, a result of mutated Tau proteins.
Image credit |
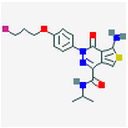
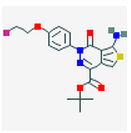
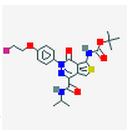
7 Figures. Seven drugs that are known to bind to neurofibillary tangles, a possible result of MAPT tau mutations. The images were found through PubChem.
Discussion
No information could be determined about the Tau proteins found in FTD, but a possible phenotype of FTD, the neurofibrillary tangles, could be researched further and possibly treated through several known drugs. Further research should be done to find small molecules that can affect the tau proteins. Both wild-type and altered tau proteins should be studied based on normal or mutated MAPT genes. Further research on using Chemical genetics in FTD research is in the Specific Aims portion of the website.
No information could be determined about the Tau proteins found in FTD, but a possible phenotype of FTD, the neurofibrillary tangles, could be researched further and possibly treated through several known drugs. Further research should be done to find small molecules that can affect the tau proteins. Both wild-type and altered tau proteins should be studied based on normal or mutated MAPT genes. Further research on using Chemical genetics in FTD research is in the Specific Aims portion of the website.
[1] Stockwell BR. Exploring biology with small molecules. Nature 2004. Dec. 16; 432 (7019): 846-54.
Site built by Kassandra Ford
Genetics 564, Spring 2015
University of Wisconsin-Madison
Site last updated: 5/13/2015
Genetics 564, Spring 2015
University of Wisconsin-Madison
Site last updated: 5/13/2015